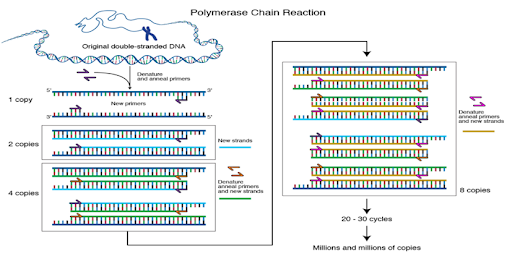
Polymerase Chain Reaction is a technique used to amplify a specific region of a DNA molecule
to generate multiple copies. This innovative technology was developed by American
biotechnologist Kary Mullis in the year 1983. He was awarded with noble prize for his innovative
work in 1993 (Singh et al., 2014). The important component of PCR include: