On the most visible and exciting applications of Genomics is in ‘Prenatal Diagnosis’. All parents want a healthy child and hence are looking for ways to predict the health of the child inside the womb. The obstetricians and maternal fetal medicine specialists are equally concerned about learning about any abnormalities or risks as early as possible. Predicting the occurrence of diseases before birth can help the expecting parents and the clinicians take more informed decisions and get a better handle on how to manage the neonate.
Discovery of any new disorder and its genetic basis immediately creates the need for predicting or preventing the recurrence of the disorder in the next generation.
A in last few years. Each technology is providing a better or a different handle to understand the genetic link to various disorders and conditions. Invariably, the application of most of these techniques is further extend to ‘Prenatal Diagnosis’, given the interest and concerns of the patients and clinicians.

Hence the ‘Prenatal Diagnosis’ space has seen the advent of a wide spectrum of technologies. While traditional high risk/instance disorders like Downs, Di-George, Cystic Fibrosis, Thalassemia etc., continue to remain the main are of interest, the advancement in sequencing technologies have increased the options for diagnosis of several other conditions which may be having a relatively lower instance rate but are equally or more debilitating.
The focus of all the technologies is to increase their accuracy (often measured in terms of specificity and sensitivity of the test) and the coverage – i.e. the variety of disorders or conditions covered by the particular test or technology.
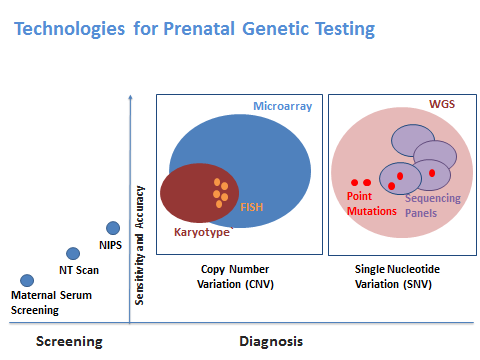
Screening vs. Diagnostic TestsBroadly, the tests presently available can be classified into ‘Screening’ and ‘Diagnostic Tests’. The former helps classify a patient into ‘high risk’ or ‘low risk’; whereas the latter helps in accurately diagnosing the presence or absence of particular conditions.
Most diagnostic tests require an ‘invasive’ procedure which in turn requires specialists for doing the procedure, the patients’ consent and counselling and the management of the patient’s concerns and risks with respect to the procedure. Due to such, costs, skill and attitude considerations, not all patients can be subject to diagnostic tests.
Hence simpler and non-invasive techniques are employed to classify the patients as low risk and high risk. Accordingly, only high risks patients are offered diagnostic procedure.

The most common and traditional tests available for prenatal screening are the biochemical markers (B-Hcg, PAAP-A, Estriol) also popularly referred to as Dual Marker, Triple Marker and Quadruple Marker based on the number of marker covered.
The advancements in Ultrasound technology and skills have provided additional methods to clinicians for identifying common genetic abnormalities (like Downs) early in the pregnancy through ‘Nuchal translucency (NT) Scan’.
Together the biochemical screening and NT scan can predict the down syndrome risk very accurately (with 95% confidence2).
The ‘Non Invasive Prenatal Screening or Testing’ (NIPS or NIPT) options have further increased the accuracy for screening of common genetic abnormalities. Most NIPS/NIPT tests have close to 99% sensitivity and specificity with respect to Trisomy 21,18,13 and few other micro-deletion syndromes.
A negative result in a screening test can provide a great assurance and relief to the expecting parents. On the other hand, a positive result in screening test should encourage them to take up a diagnostic test.
Poor understanding of the variety of genetic abnormalities and a blind faith or overdependence on the screening techniques are risk factors which need to be addressed through a more conscious and aware practice in the fetal medicine arena.
From Screening to DiagnosisOnce a pregnancy has been classified as high risk based on any of the screening tests or due to other factors such as age, previous child etc., the parents are offered diagnostic tests. The diagnostic test often involves an invasive procedure such as Amniocentesis or Chorionic Villus Sampling to take out a sample of the cells belonging to the fetus. Doing a chromosomal analysis or a molecular study on the fetal tissue can provide a definitive diagnosis of the disorder.
The traditional chromosomal analysis techniques like Karyotyping or Fluorescent In Situ Hybridization (FISH) are very popular for a definitive diagnosis of the common disorders.
Beyond Common Genetic AbnormalitiesSo far, the larger concern in Prenatal Diagnosis has been with respect to genetic disorders which are relatively common (higher instance rate in a population) like Down Syndrome. However, the combined probability of all other genetic syndromes including the single gene defects and the various micro-deletion/duplication syndromes presents and equally large if not a larger challenge for the parents. Here the role of carrier screeningalso become important in order to again classify the pregnancy as high risk depending on the carrier status of both the parents. When the suspicion is with respect to a single gene disorder or point mutation the various molecular techniques such as Polymerase Chain Reaction (PCR), MLPA (Multiplex Ligation-dependent Probe Amplification), and Sanger Sequencing can help diagnosis or rule out the suspected disorder or mutation.
Whole Genome Approach: CMA and NGSIn cases where there is no single mutation which is suspected or the risks pertain to a wider set of mutations; there is a need for a whole genome approach. In this approach, the entire genome is scanned for identification of variations or genetic changes.
Newer technologies such as Chromosomal Microarray Analysis, also commonly known as Microarray provide a ‘whole genome approach’ for detection of ‘Copy Number Variations’. Microarray is also useful in unearthing Loss of Hetrozygosity (LOH) and micro-deletions and micro-duplications which are not detected by the traditional chromosomal analysis techniques. Microarrays can also be used in cases where there is little or no information about the abnormalities in the previous child or pregnancy. Microarrays have now become more common in prenatal diagnosis and have been endorsed as the appropriate test by American College of Obstetricians and Gynaecologists (ACOG) and Society for Maternal Fetal Medicine (SMFM)
Similarly, the Whole Exome Sequencing OR Whole Genome Sequencing can help in dealing with Single Nucleotide Variations (SNV) using the Next Generation Sequencing (NGS) technology. Given that the number of SNVs identified in any patient can be very large and may include novel variations as well, it is critical to know what kind of variation we want to confirm or rule out. Hence testing of previous child and parents becomes critical when we use the NGS technologies for prenatal testing.

As is evident, there are a wide variety of technologies available for prenatal screening and diagnosis. Each technology has its own advantage and applications. Newer technologies like Microarray and NGS though more comprehensive are equally more complex in terms of interpretation and reporting.
It is important for the clinicians to be more aware about the uses and limitations of various techniques and to consult with geneticists, genetic counsellors and genetic laboratories to identify the most appropriate technique for different cases.
References:-
//getthediagnosis.org/definitions.html#sthash.dzn7xcCb.dpuf.
- Nicolaides KH , Screening for fetal aneuploidies at 11 to 13 weeks, Prenat Diagn 2011; 31: 7–15,
- Committee Opinion #581 “The Use of Chromosomal Microarray Analysis in Prenatal Diagnosis” is published in the December issue of Obstetrics & Gynecology.

